THURSDAY, 1 OCTOBER 2009
Antibodies are an astonishing product of the immune system. These Y-shaped proteins, which are made by a subset of white blood cells, are central to the destruction and removal of infectious diseases.The function of an antibody is simple: to recognise and stick to a specific molecular ‘label’ – called an antigen – on the surface of an unwelcome visitor in the body. These pathogens, which may be bacteria, viruses or parasites, are unlike human cells and each pathogen will have its own unique antigen. Once antibodies are stuck to the pathogen, it can usually be destroyed or neutralised by other components of the immune system.
Does this mean that our genes carry the code for every antibody we may ever need, to deal with any of the countless pathogens that we may encounter during our lives?
The answer is no, because the immune system has developed a far better way to generate the antibodies it needs: evolution on a molecular scale. This evolution, as we might expect, occurs through genetic mutation and promotes survival of the fittest – but the process takes weeks rather than centuries. The result is the expression of strongly binding, highly specific antibodies against any pathogen that has been detected.
The first ingenious part of this antibody evolution comes from the requirement for diversity. For the most strongly-binding antibodies to be preferentially selected, there has to be a massive range of antibodies to choose from, each with a slightly different antigen shape preference.
B lymphocytes (B cells), the cells that make antibodies, are indeed created in a way that means each expresses one of an enormous repertoire of antibodies. This is achieved by the unique design of the DNA sequences that code for each different antibody protein.
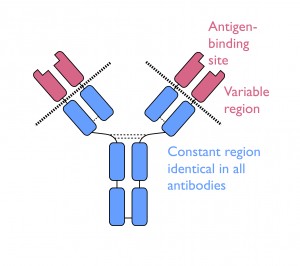
The DNA sequence for every antibody is identical except for the small part that determines the antigen-binding region found at the tips of each Y-shaped protein molecule. This section, called the ‘VDJ’ unit, is formed by selecting three units from three different regions along the gene (called V, D and J). In humans, there are around 65 V units, 27 D units and 6 J units and any combination is possible – a sort of genetic pick ‘n’ mix. But that’s not all; the complete antigen-binding site is constructed from two separate protein chains whose antigen-binding regions are coded for by different VDJ units – in other words, a combination of two already random pick ‘n’ mix selections! The number of possible sequence combinations is enormous. It is estimated that B cells express as many as 100 billion different antibodies, with each cell rearranging its DNA to randomly encode one unique antibody with its own unique binding site.
Following an infection, a selection process takes place. The B cell initially expresses its antibodies on its surface where they act as receptors. This allows the cell to first find out whether its antibody is of any use, by seeing if it will recognise any of the pathogenic antigens. This initial step takes place in the spleen and also in the lymph nodes.
Antigens from invading pathogens are carried by cells, conveniently named antigen-presenting cells (APCs), from infection sites to these locations. APCs display antigens to the antibody receptors on vast numbers of B cells, in the hope that a small number have antibodies with a good fit.
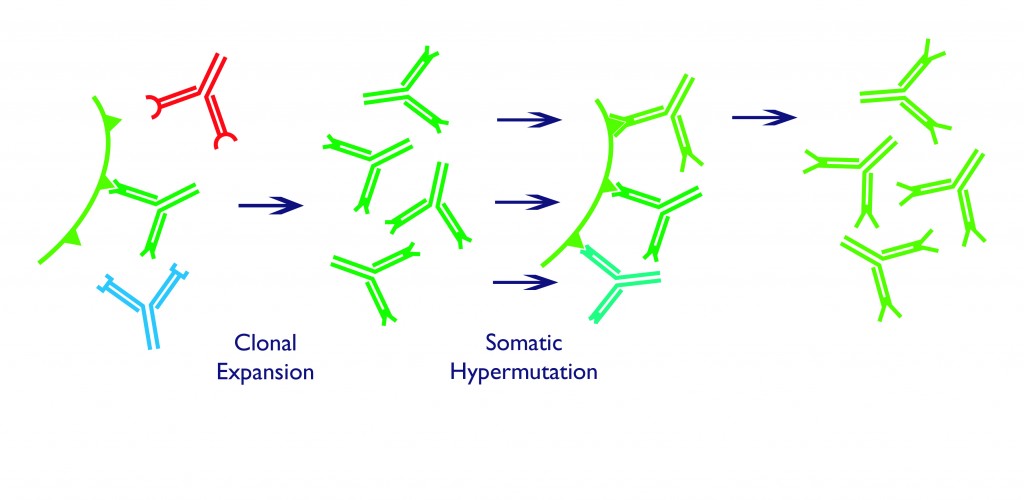
Those B cells fortunate enough to meet this requirement receive an activating signal from another type of lymphocyte, called a T cell. These signals promote the survival, growth and division of the B cells to create more copies of only those that express the correct antibody in a process called clonal expansion. The army of activated, antigen-specific B cell clones forms a cluster called a germinal centre, where they use a further trick to generate even better fitting antibodies.
Somatic hypermutation occurs in germinal centre B cells that have already proven their worth by activation upon binding to an antigen. Within these cells, the activation initiates a process in which the DNA is broken in two, at the V regions described above. The join is then repaired, but in an error-prone way, which leads to point mutations in the DNA sequence. These tiny sequence changes alter the shape of the antigen-binding region in the encoded antibody – some changes improve binding, while others make the antibody worse than it was before.
The thought of mutation occurring in our bodies probably brings about images of cancer or deformities – far from desirable consequences. Yet evolution has encouraged high mutation rates in B cells to provoke a rapid natural selection process.
B cells fight amongst themselves in a competition for survival signals, with the lottery of somatic hypermutation adding even more variation to their antibody repertoires. The few whose antibodies have been improved by this process are preferentially selected. Only the fittest cells survive, and in just a couple of weeks after an infection, this induced natural selection process results in long-living B cells which secrete their specific antibodies into our blood, sometimes for the rest of our lives.
It is this antibody generation that provides us with life-long protection against potentially life-threatening pathogens and is the process that is stimulated by vaccinations. It is amazing to think that the process of evolution through natural selection not only underlies the adaptation of species to their environments, but is also the mechanism used by our immune system to keep us alive.
Robert Williams is a PhD student in the Babraham Institute.